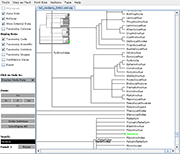
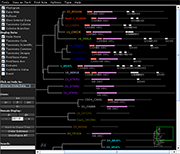
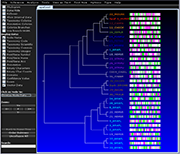
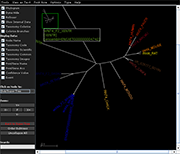
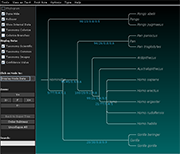
Archaeopteryx is a software tool for the visualization, analysis, and editing of
potentially large and highly annotated phylogenetic trees.
Archaeopteryx (the successor to ATV)
is entirely written in the Java programming language (it is based on the forester libraries). It can be used both
as applet (ArchaeopteryxA and
ArchaeopteryxE) and as a standalone application.
Documentation and answers for some frequently asked questions are (will be) available
here
(some older documentation is also available
here),
phyloXML related issues might be discussed here.

|

|

|

|

|

|

|

|

|
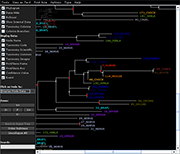
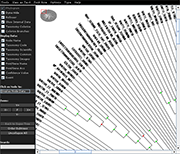
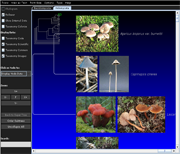
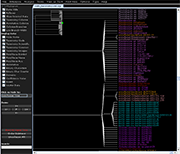
| Species colorization and gene duplication display (demo) | Circular display type (demo) | Species images (demo) | Large trees (proteobacteria from NCBI taxonomy) | Search function (demo) | Protein architecture display (demo) | Expression value display | Unrooted display type (demo) | Multiple support values (demo) |
Archaeopteryx requires Sun Java version 5 (also
called Java 1.5) or higher. Archaeopteryx applets (ArchaeopteryxA, ArchaeopteryxE) require
the Sun Java plug-in. Most web browsers under Windows and Apple OS X have this
already installed. Under Linux, this plug-in might need to be installed, though.
For example, under Ubuntu Linux, the Sun Java plug-in can be downloaded and
installed with the following command: "sudo apt-get install sun-java6-jre
sun-java6-plugin". More information is available here.
Archaeopteryx has been tested on Windows, Linux, and Apple Macintosh (OS X with
Java 1.5.0_13).
Latest version: Archaeopteryx 0.968 beta BG (2012.01.03)
Jar file containing Archaeopteryx: » forester.jar
To download, right-click and then select "Save Link As..." (on Windows), then
launch Archaeopteryx by clicking the downloaded "forester.jar" file on your own
desktop.
Example configuration file: » _aptx_configuration_file
Jar file containing only Archaeopteryx applets (only needed for web developers): »
archaeopteryx_applets.jar
Sign up for the Archaeopteryx mailing list to be
informed of new versions and changes.
Previous versions
ArchaeopteryxA examples (open own frame)
Answers for some frequently asked questions are (will be) available at aptxevo.
Clicking on the "forester.jar" file should start it, the configuration file ("_aptx_configuration_file") will be used if it is in the same directory as the jar file.
command line (without use of a configuration file or "_aptx_configuration_file" is
in the same directory as the jar file):
"java -cp path\to\forester.jar
org.forester.archaeopteryx.Archaeopteryx"
command line for using a configuration file anywhere in the file system:
"java -cp path\to\forester.jar org.forester.archaeopteryx.Archaeopteryx -c
path\to\_aptx_configuration_file"
command line for directly opening a treefile and using a configuration file
anywhere in the file system:
"java -cp path\to\forester.jar org.forester.archaeopteryx.Archaeopteryx -c
path\to\_aptx_configuration_file treefile"
Since the Java default memory allocation is quite small, it might by necessary (for
trees with more than around 5000 external nodes) to increase the memory which Java
can use, with the "-Xms" and "-Xmx" Java command line options.
For example:
"java -Xms32m -Xmx256m -cp path\to\forester.jar
org.forester.archaeopteryx.Archaeopteryx -c
path\to\_aptx_configuration_file"
To avoid typing, it is easiest to create at batch (.bat) file. For example create a
new file named "aptx.bat" and put a line like this into it:
java -Xms32m -Xmx512m -cp "C:\path\to\forester.jar"
org.forester.archaeopteryx.Archaeopteryx -c
"C:\path\to\_aptx_configuration_file"
Clicking on "aptx.bat" should now start Archaeopteryx.
A description about how to use Archaeopteryx as Java applet in your own website is available here.
Certain programs produce New Hampshire formatted trees (possibly
inside a Nexus file) where internal tree nodes have both branch length as well as
support values [e.g. "(a:0.1,b:0.2)0.90:0.1"]. By default, Archaeopteryx
interprets these support values (the "0.90" in the example) as node names. In most
situations this is not a big problem, except if the trees are to be rerooted --
then the "support values" move to the wrong branches (because node names are
attached to a node and not to a branch like support values).
To correctly interpret such internal numbers as support values, check "Internal
Numbers Are Confidence Values" under the "Options" menu (in the Archaeopteryx
application) before reading the tree file.
The source code for Archaeopteryx (as application/module of the the open source forester libraries) is available on Google code: //code.google.com/p/forester/
Please consider joining the Archaeopteryx mailing list to
be informed of updates and changes
(probably no more than about one message per month).
Please let us know if you have a suggestion for this list.
Han M.V. and Zmasek C.M. (2009). phyloXML: XML for evolutionary
biology and comparative genomics. BMC Bioinformatics, 10:356. [PubMed] [BMC Bioinformatics] [PDF]
Later...
2026 © Copyright www.phylosoft.org | All Rights Reserved